|
|
| Untersuchte Arbeit: Seite: 7, Zeilen: 1-13 |
Quelle: Abate-Shen 2002 Seite(n): 780, Zeilen: figures |
|---|---|

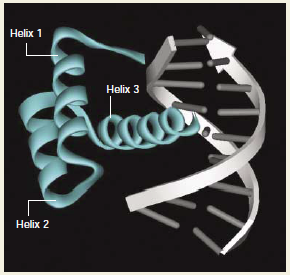
Figure 4. The structure of the homeodomain. Despite its considerable variation in primary sequence, the three-dimensional structure of the homeodomain has been well conserved, and corresponds to three α helices that have a relatively unstructured amino-terminal arm. The consensus amino acids, which are invariant among all homeodomains, are essential in maintaining structural integrity and making essential contacts with DNA. Amino acids that distinguish the various homeoprotein families contribute to differences in their DNA-binding specificities and other functional properties. The figure shows the Engrailed–DNA complex (Figure adapted from Abate-Shen, 2002) Figure 5. Generating diversity among homeoproteins. The distinct classes of homeoproteins show considerable sequence diversity within the homeodomain (top), and within the amino- and carboxy-terminal regions that flank this domain (bottom). The sequence diversity is believed to contribute to their distinct functional properties, by generating distinct DNA-binding specificities, promoting unique protein–protein interactions, and other mechanisms. [Functional diversity is also generated by additional conserved domains or motifs that occur in particular homeoprotein families. HOX proteins contain a hexapeptide (HP) motif-a short stretch of conserved residues that are responsible for mediating interactions with PBX homeoproteins. MSX proteins also have short stretches of conserved amino acids that flank the homeodomain, called the extended homeodomain (EHD), although their functions are unknown. PAX proteins contain an additional DNA-binding domain, known as the paired box. Members of the SIX family have a conserved amino-terminal domain called the Six domain, and LIM homeoproteins are named after a protein interaction motif, the LIM domain (Figure adapted from Abate-Shen, 2002).] |
Box 3 Generating diversity among homeoproteins
The distinct classes of homeoproteins show considerable sequence diversity within the homeodomains (top), and within the amino- and carboxy-terminal regions that flank this domain (bottom). The sequence diversity is believed to contribute to their distinct functional properties, by generating distinct DNA-binding specificities, promoting unique protein–protein interactions, and other mechanisms. Functional diversity is also generated by additional conserved domains or motifs that occur in particular homeoprotein families. HOX proteins contain a hexapeptide (HP) motif-a short stretch of conserved residues that are responsible for mediating interactions with PBX homeoproteins. MSX proteins also have short stretches of conserved amino acids that flank the homeodomain, called the extended homeodomain (EHD), although their functions are unknown. PAX proteins contain an additional DNA-binding domain, known as the paired box. Members of the SIX family have a conserved amino-terminal domain called the Six domain, and LIM homeoproteins are named after the protein interaction motif, the LIM domain.
Despite its considerable variation in primary sequence, the three-dimensional structure of the homeodomain has been well conserved, and corresponds to three α-helices that have a relatively unstructured amino-terminal arm. The consensus amino acids, which are invariant among all homeodomains, are essential in maintaining structural integrity and making essential contacts with DNA. Amino acids that distinguish the various homeoprotein families contribute to differences in their DNA-binding specificities and other functional properties. The figure shows the Engrailed–DNA complex. |
The source is given at the end of the two figure captions. However, both figures are copied and not "adapted". Furthermore, the extensive figure captions are also taken 1-to-1 from the source, which does not become clear from the references at all. |
|